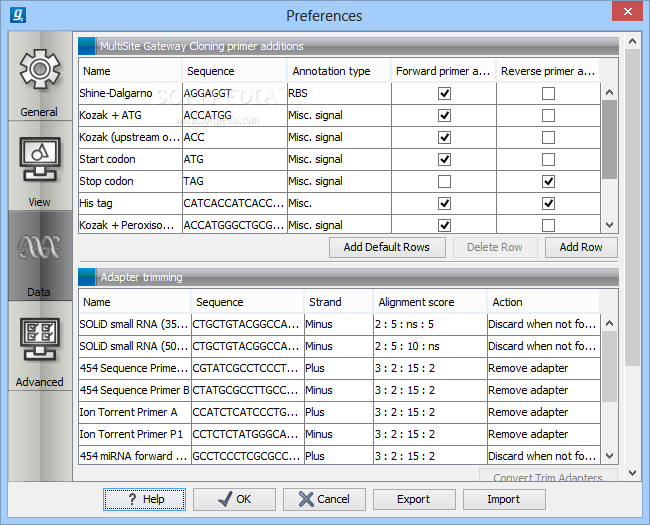
UPDATED to include a multiple sequence alignment and maximum likelihood tree workflow, and an optional direct BAM import workflow for those who don't want to first import data to CLC prior to running the workflow. However, if users have access to the bed file for this workflow, this can be included into the workflow immediately after read mapping, but adding our "Trim Primers & Dimers" tool. Due to the way TF AmpliSeq data is generated, no primer BED file is provided. This zip contains an implementation of a CLC Genomics Workbench workflow suitable for use with ThermoFisher Ion AmpliSeq SARSCOV2 panel data. Also, position specific primer trimming, primer/dimers, and chimera filtering is included, similar to Nick Loman's pipeline, but the results may nonetheless have some differences. Variant calling is done with CLC using our Low Frequency Variant Calling with relatively leanient settings "inspired by" those in the Medaka pipeline, but care should be taken in evaluating variants or quasispecies results. The nanopore workflows use an implemetation in CLC of minimap2/racon included with CLC Genomics Workbench - and not Medaka, which is being used by several other labs. This zip archive includes workflows intended for both Illumina and Nanopore data produced from the ARTICv3 protocol. There many other ways to build trees in CLC Genomics Workbench for this purpose, and other "good" ways to build SARS-CoV-2 phylo trees would be to use a NJ tree as a starting guide to then run maximum likelihood estimation tree and/or use read mapping data to build SNP based trees of all samples. The workflows include a basic qc/mapping/consensus calling workflow an advanced workflow that extends the basic workflow to include indel, structural variant calling, local realignment of potential indel regions, and a low frequency variant detection step and a simple fast multiple sequence alignment and neighbor-joining tree building workflow. This zip file contains three workflows for CLC Genomics Workbench that are designed to work with QIAGEN's QIAseq SARSCOV2 amplicon sequencing panel. The zip archive will be automatically unzipped and the workflows can be open, edited, run directly, or installed for other users on the same system. zip archives provides above and import them into CLC Genomics Workbench using the Import->Standard Import option on the tool bar. Please contact me via email ( or Twitter ( if there's any major issue with any of these pipelines that need to be fixed. 
To the best of my ability, I have created these workflows to take into account aspects of the wet-lab protocols (where ever possible), and may include specific primer sequences, adapter sequences, or other parameter settings to account for differences in sequencing platforms(i.e. Each workflow includes - a minimum - qc, trim, mapping, variant calling and the output of a consensus sequence. The zip archives available here are intended to provide CLC Genomics Workbench users with a quick way to get started with analysis of SARS-CoV-2 genomics data from a variety of upstream laboratory protocols. CLC Genomics Workbench Workflows for SARS-CoV-2 Background QIAGEN kit handbooks and user manuals are available at ( or can be requested from QIAGEN Technical Services or your local distributor. For up-to-date licensing information and product-specific disclaimers, see the respective QIAGEN kit handbook or user manual. These products are not intended for the diagnosis, prevention or treatment of a disease. **ĭISCLAIMER QIAGEN Digital Insights provided here are intended for molecular biology applications only. If you have any questions - please DM me on Twitter or here on GitHub. They are intended as examples in the hopes that users may find them useful to get started quickly on analyzing their SARS-CoV-2 genomics data with CLC. ** The workflows found here are provided AS IS by Jonathan Jacobs, and not officially by QIAGEN Digital Insights. Please visit our COVID-19 Emergency Response Licenses page to request your licenses and access additional resources. Have fun!Īlso, free Extended Trials of both CLC Genomics ProSuite and Ingenuity Pathway Analysis are available from QIAGEN Digital Insights until June 15th, 2020. There you will find some downloadable workflows that can be imported, edited, and run in CLC. If you are looking for some example workflows for using CLC with SARSCOV2 data - look under the "CLC-SARSCOV2-Workflows" folder. 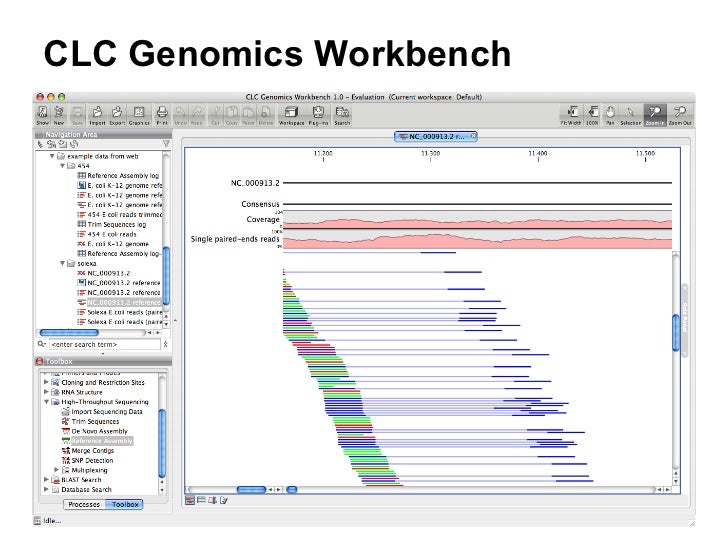
An unofficial repo for CLC Genomics Workflows, custom datasets, tutorials, and other files.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
